I’m an incoming Tenure-Track Assistant Professor at the Department of Computer Science and Engineering at the Hong Kong University of Science and Technology, starting Fall 2026. I’m currently a NIH K99/R00 Postdoctoral Fellow at the Department of Biomedical Informatics at Harvard Medical School, working with Nils Gehlenborg. I received my PhD in Computer Science and Engineering from Seoul National University with the supervision of Jinwook Seo at the Human-Computer Interaction Lab.
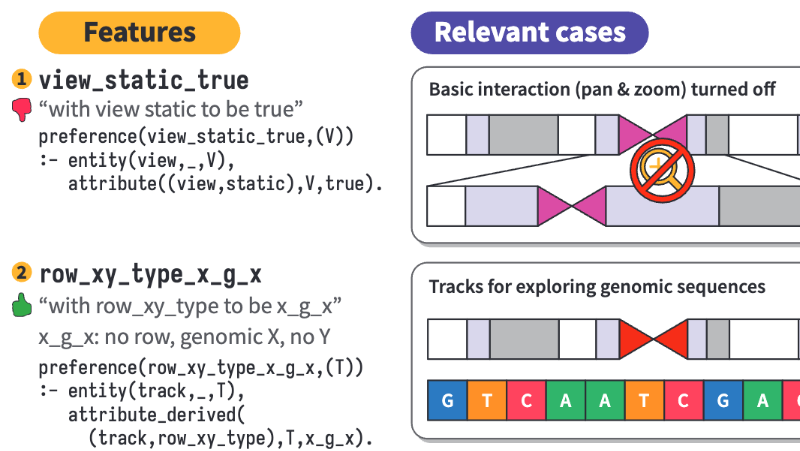
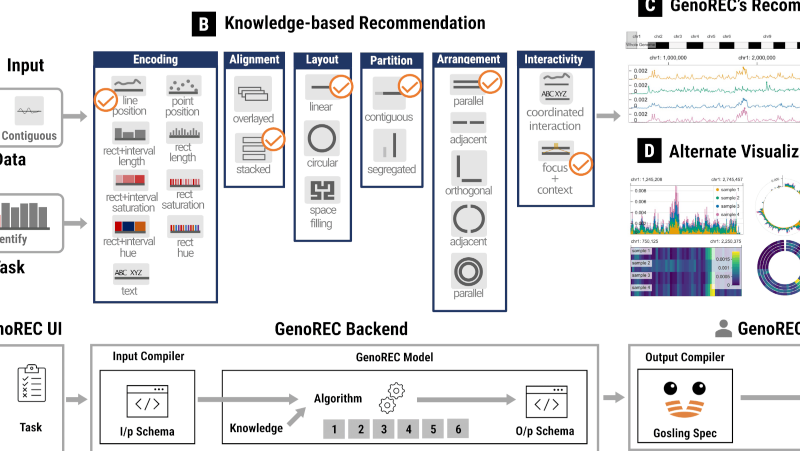
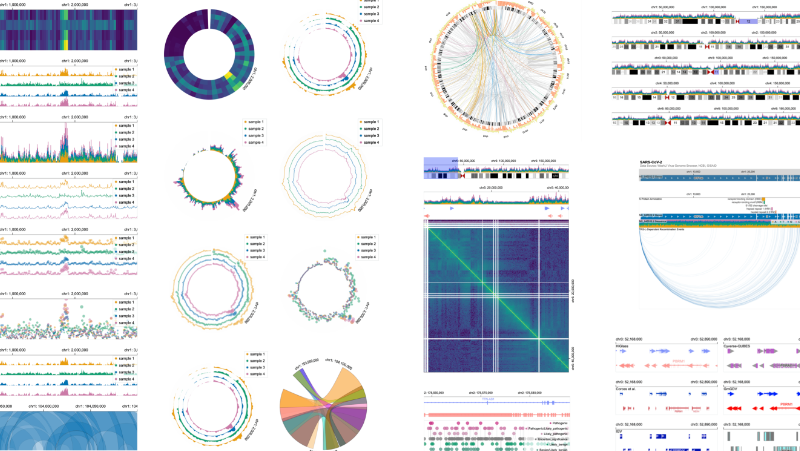
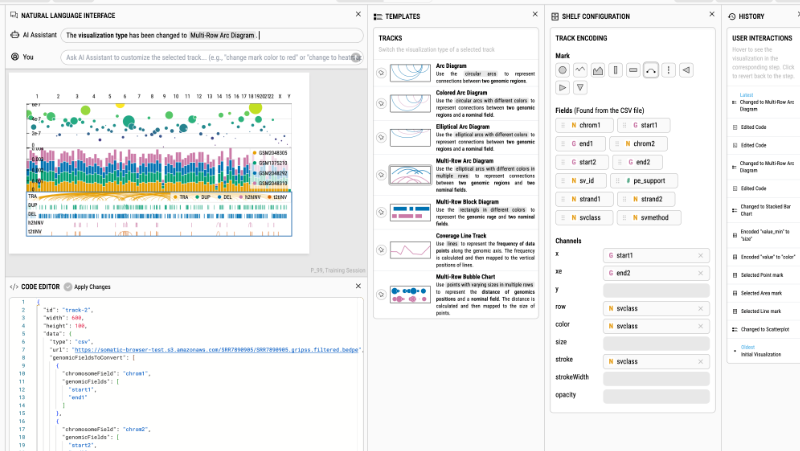
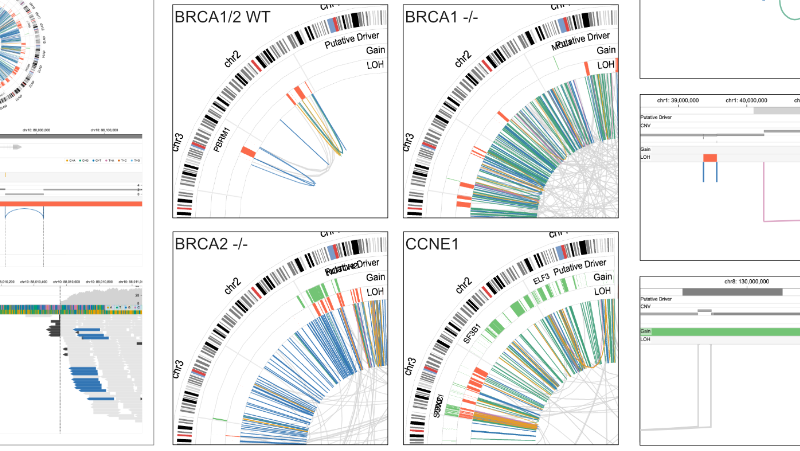
As an interdisciplinary researcher, I work at the intersection of visualization, human–AI interaction, and biomedicine. I have designed and developed interactive data systems that scale to a wide range of user needs within biomedical domains. For example, the Gosling visualization grammar enables constructing scalable and interactive genomics data visualization and leveraging AI methods (read Nature’s Technology Feature).
My work has been published in top-tier conferences and journals across my fields, including VIS, TVCG, CHI, UIST, Nature Methods, Bioinformatics, and npj Digital Medicine. My work has been recognized by academic awards, such as NIH/NHGRI K99/R00 Pathway to Independence Award, VIS Best Paper Honorable Mention Award, ISMB BioVis Best Abstract Award, ISMB BioVis Runner-Up Abstract Award, and VIS Best Poster Research Award. I have been serving as a program committee member for top conferences in visualization, including VIS, EuroVis, and PacificVis.
News [see all]
Selected Projects
Selected Publications [see more]
Media Coverage
 Nature (TECHNOLOGY FEATURE)
Nature (TECHNOLOGY FEATURE)
Powerful 'grammar' allows geneticists to display their data in interactive and scalable illustrations.
"Postdoc Sehi L'Yi, who led Gosling's development, says that what differentiates Gosling from other visualization tools is its expressiveness. With most tools, he says, the graphics that can be made and what they will look like are predefined. 'It is really not easy to customize visualizations as a user.' But with Gosling, users can, for instance, specify the colour, dimensions and placement of the symbol used to represent a centromere or genomic interval, then overlay that on an ideogram of a chromosome to highlight a region of interest."
|

|